MRI Analyze Roots Tool¶
The tool allows to analyze images of small roots. It measures the length and area of the main root, the number of secondary roots and the length and area of the whole root.
Getting started¶
To install the tool, save Tracing_.jar in the plugins folder of your FIJI-installation. Drag the link MRI_Analyze_Roots_Tool.ijm to the ImageJ launcher window, save it under macros/toolsets in the ImageJ installation and restart ImageJ.
Select the "MRI_Analyze_Roots_Tool" toolset from the >> button of the ImageJ launcher.

- the first button (the one with the image) opens this help page
- the a-button analyzes the current image
- the b-button runs the batch analysis on a folder containing the input images
Options¶
By right-clicking on the a-button, you can open the options dialog.
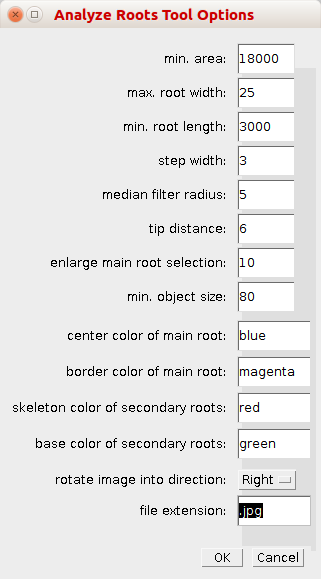
- min. area: Objects smaller than min. area pixel will not be considered roots and will be excluded from the analysis
- max. root width: parameter for the tracing algorithm - the maximum diameter of the root
- min. root length: parameter for the tracing algorithm - the minimal length of a root
- step width: parameter for the tracing algorithm - the step width used by the tracing to find the next point on the root
- median filter radius: radius of the median filter applied before the tracing
- tip distance: distance from the lowest point of the root's tip to the point where the tracing begins
- enlarge main root selection: Number of pixel by which the selection of the main root is enlarged to detect second order roots
- min. object size: the minimum size of second order roots within the enlarged selection of the main root
- center color of main root: the color of the center of the tracing of the main root
- border color of main root: the color of the border of the tracing of the main root
- skeleton color of secondary roots: the color of the skeleton of the secondary roots
- base color of secondary roots: the color of the bases of the secondary roots
- rotate image into direction: rotate the image to the right or to the left or not at all before the analysis
- file extension: the file extension of the input files
Results¶
When used on a single image (a-button), a results table with the following measurements will be created:
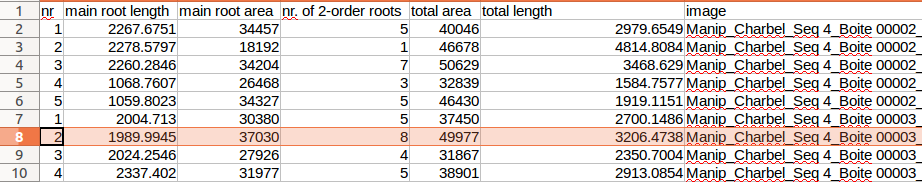
- main root length - the length of the central line of the tracing of the main root
- main root area - the surface of the tracing of the main root as determined by the right and left border lines
- nr of second order roots - the number of 2. order roots - it is determined by removing the main root, enlarging the tracing of the main root and counting the objects found within
- total area - the bounding box of each root is determined, a threshold is applied to the region within the bounding box and the number of object pixels is counted
- total length - The main root is removed from the image, a threshold is applied to the remaining signal and the resulting binary mask is skeletonized - the macro Measure Skeleton Length is used to measure the total length and the length of the main root is added to the result
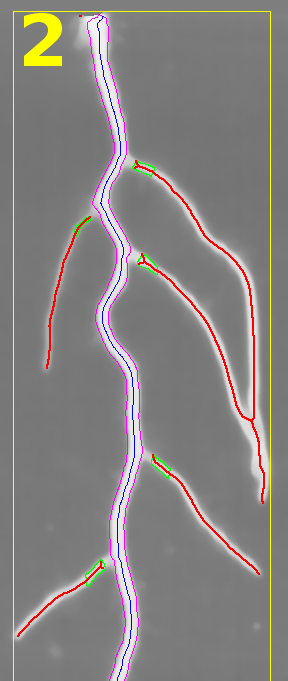
The results found by the tool are displayed in an overlay on the input image.
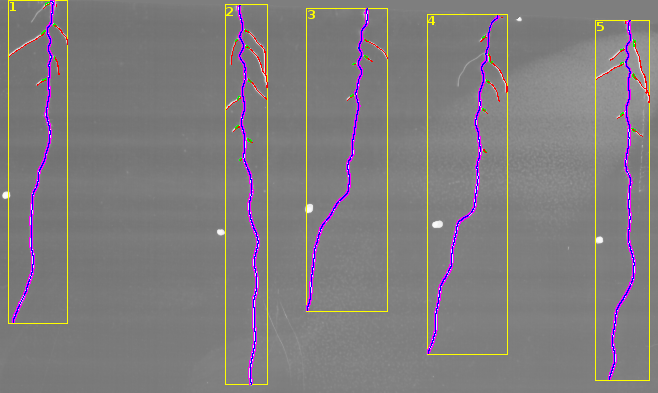

- yellow - the bounding box of the root and its number in the image that corresponds to the number in the measurements
- blue - the central line of the tracing of the main root
- magenta - the border lines of the tracing of the main root
- green - the parts of the second order roots near the main root that are counted
- red - the skeletons of the second order roots that are used to measure the total length
Methods used¶
The tracing is an implementation of the algorithm described in [Can99]. Once the tracing is done, the outer borders are connected to form an area selection. The area selection is enlarged by some pixel and the content removed. It is then enlarged again and the number of objects within are counted as 2. order roots.
- Ali Can, Hong Shen, Turner, Tanenbaum, and Roysam (1999). Rapid automated tracing and feature extraction from retinal fundus images using direct exploratory algorithms. IEEE Transactions on Information Technology in Biomedicine 3, 125–138.
After the main root has been removed the remaining signal in the bounding box of the root is thresholded and the resulting mask is skeletonized. The length of the skeletons is measured using an implementation of [Nie05]
- Niemisto, A., Dunmire, V., Yli-Harja, O., Wei Zhang, and Shmulevich, I. (2005). Robust quantification of in vitro angiogenesis through image analysis. IEEE Transactions on Medical Imaging 24, 549–553.